Previously, we wrote about the unique capabilities that next-generation sequencing (NGS) offers the oncology clinic. NGS could mark the beginning of a shift away from “single-site” technologies such as FISH and PCR-based testing, in favor of comprehensive screening across many different targets at once.
One prerequisite for replacement of these legacy methods with NGS is for clinical labs to demonstrate equivalent or superior performance relative to current standard-of-care tests. Since many different types of mutations are known to be important for formation, classification, outcome prediction, and treatment of myeloid cancers, labs will be challenged to generate very comprehensive data in support of NGS-based clinical testing. Consider the following categories of variants for which robust analytic performance will need to be both established and maintained.
Diagnostic mutations
Distinguishing between different types of cancers relies on very specific criteria that are developed by groups such as the World Health Organization and National Comprehensive Cancer Network. Correct diagnosis is important to ensure patients receive optimum care for the particular disease in question. In addition to parameters such as cell counts and morphology, the presence of mutations in certain genes is being increasingly used to aid pathologists in making definitive diagnoses.
For example, JAK2 mutation is included in the WHO diagnostic guidelines for polycythemia vera (PV), a slow-growing malignancy in which the bone marrow produces an overabundance of erythrocytes. Initially, the point mutation JAK2 V617F was estimated at 97% of cases of PV; however, this mutation may also be present in other myeloid malignancies. In addition, other mutations in JAK2 have since been characterized for PV, such as those affecting exon 12. On the other hand, in addition to JAK2, mutations in the calreticulin gene (CALR) or the myeloproliferative leukemia virus oncogene (MPL) are associated with essential thrombocythemia and primary myelofibrosis, but rarely PV11.
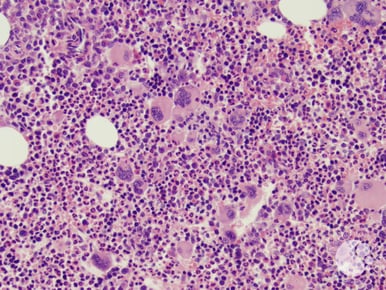 Hematoxylin and eosin stain of bone marrow core biopsy showing polycythemia vera.
Hematoxylin and eosin stain of bone marrow core biopsy showing polycythemia vera.
Image courtesy of American Society of Hematology image bank
In short, diagnosis of myeloid cancers is very complex. And while comprehensive genetic screening can tremendously enhance clinicians’ ability to differentiate disease, the ever-increasing number of relevant targets will necessitate equally comprehensive due diligence to ensure these tests are performing as expected.
Prognostic markers
In the context of molecular genetics, prognostic markers are mutations that are correlated with a particular course of disease. They play a key role in determining treatment since patients who have a variant associated with poor outcome should be considered for a regimen equipped to combat more aggressive forms of the disease.
One example of a putative prognostic signature for myeloid malignancies is mutations in the gene ASXL1. The ASXL1 protein regulates gene expression via complex mechanisms, and mutations that alter its function may be correlated with adverse events such as disease progression. Duplication of a guanine at cDNA position 1934, which results in the frameshift G646fs*12, accounts for over half of all ASXL1 mutations. Interestingly, reliable detection of this mutation has proven challenging due to low-complexity sequence around the variant, and there have been conflicting reports as to whether this mutation is real, or a technical artifact. Thus, this variant highlights both analytical and clinical challenges that must be overcome prior to a physician gaining information that is useful in clinical practice2.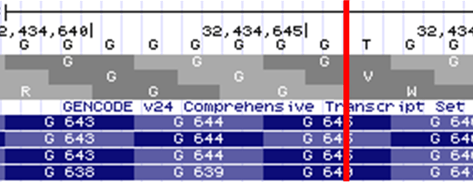 
The human genome browser at UCSC. Genome Res. 2002 Jun;12(6):996-1006.
Image courtesy of Kent WJ, Sugnet CW, Furey TS, Roskin KM, Pringle TH, Zahler AM, Haussler D.
USCS genome browser zoomed view of ASXL1. GRCh38 coordinates are at the top, above the nucleotide track. The location of G646fs*12, between chr20:32,434,646-32,434,647 is indicated by the red bar. Note the poly-G track present at this location.
Therapeutic indications
Perhaps one of the most well-known successes thus far in precision medicine is the story of imatinib mesylate for chronic myeloid leukemia (CML). Approved by the US FDA in 2001 under the trade name Gleevec® (Novartis), imatinib has gained a reputation as a “silver bullet” drug, having shown extremely high efficacy even for late-stage CML patients.
Over 95% of cases of CML are caused by chromosomal translocation that results in fusion of the BCR and ABL1 genes to form BCR-ABL1, also known as the Philadelphia chromosome (this fusion gene was first described by researchers from Fox Chase Cancer Center and U Penn School of Medicine, both Philadelphia-based). BCR-ABL1 drives cancer formation through increased kinase activity, which leads to dysregulation of signaling pathways that control cell proliferation and survival. Imatinib, a member of a class of drugs known as kinase inhibitors, works to decrease the activity of the abnormal enzyme resulting from the BCR-ABL1 fusion gene, in an effort to restore cellular processes to their native, or normal, state.
Unfortunately for many CML patients who initially respond well to imatinib, there is more to the story. As is the case for many disease interventions, kinase inhibitor therapy can be viewed from an evolutionary perspective as a bottleneck, which in this case selects for clones with a fitness advantage in the presence of imatinib. Clonal evolution of CML gives rise to cells carrying imatinib-resistance mutations such as ABL1 T315I. In turn, new therapies have been developed such as ponatinib, a generation III kinase inhibitor that was initially approved by the FDA in 2012 (Iclusig®, Ariad), and is intended for Ph+ CML (or acute lymphoblastic leukemia) patients who have become resistant to prior tyrosine kinase inhibitor therapy.
Beyond targeted therapies for BCR-ABL1, there have been many promising advances in medicines for myeloid cancers – a few noteworthy examples from around the time of publication of this article are summarized below.
Fms-like tyrosine kinase 3 (FLT3) plays a role in survival, proliferation, and differentiation of hematopoietic stem cells. Mutations in this gene are frequently found in acute myeloid leukemia (AML), and include a unique mutation type known as an internal tandem duplication (ITD) wherein a particular sequence is duplicated, and the duplication is present juxtaposed to the native copy. Point mutations in the tyrosine kinase domain such as FLT3 D835Y also lead to aberrant signaling and dysregulation of cellular function. Many different tyrosine kinase inhibitors are being evaluated in clinical trials to determine safety and efficacy for FLT3-mutated AML, and in April 2017, the FDA granted the first approval for a FLT3 inhibitor, midostaurin (RYDAPT®, Novartis).
USCS genome browser zoomed view of ASXL1. GRCh38 coordinates are at the top, above the nucleotide track. The location of G646fs*12, between chr20:32,434,646-32,434,647 is indicated by the red bar. Note the poly-G track present at this location.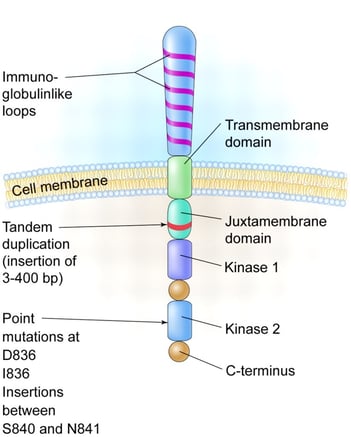
Structure of Flt3 receptor tyrosine kinase with locations of occurrence for ITDs and pathogenic point mutations. Image courtesy of Litzow M. “More flitting about FLT3.” Blood 2005 106:3331-3332, doi: https://doi.org/10.1182/blood-2005-08-3475. Accessed 3 Oct 2017.
Isocitrate dehydrogenase 1 and 2 (IDH1 and IDH2) are homologous genes whose protein products are key metabolic enzymes that convert isocitrate into α-ketoglutarate. Certain mutations in these genes result in neomorphic enzyme activity by which the metabolic product 2-hydroxyglutarate is produced. This aberrant species has been referred to as an “oncometabolite”, given its ability to disrupt cellular processes and lead to oncogenesis, particularly for formation of hematologic malignancies. In August of 2017, the FDA approved enasidenib (Idhifa®, Celgene) the first mutant IDH2 inhibitor to enter the clinic for AML.
- BRAF V600E, hairy cell leukemia, and vemurafenib7
Hairy cell leukemia (HCL) is a slow-growing malignancy that causes the bone marrow to produce too many lymphocytes. For many years, the underlying genetic cause of HCL was unknown, until it was discovered that BRAF V600E, the same point mutation that has high prevalence in melanoma, is the main driver for this rare cancer. The first BRAF inhibitor, vemurafenib (Zelboraf®, Genentech), was approved by the FDA in 2011 for unresectable metastatic melanoma; there is currently an ongoing Phase II clinical trial to evaluate the safety and efficacy of vemurafenib for HCL.
The categories of mutations presented above are not mutually exclusive - it is entirely possible for detection of a single mutation to allow a clinician to positively identify a disease, determine how aggressively and by what mechanisms it will likely progress, and to provide treatment designed to specifically target cancer cells, while leaving normal tissues (ideally) unperturbed. Whether this mutation is a translocation, an internal tandem duplication, a simple point mutation, or yet a different type of genetic alteration, the analytic performance capabilities of genetic tests are central to the success of drug development, as well as the practice of clinical oncology.
SeraCare is proud to offer solutions that will help ensure next-generation diagnostics are poised to enable precision medicine. We are pleased to begin serving the hematology/ oncology community with the latest additions to our clinical genomics portfolio, Seraseq™ Myeloid Mutation DNA Mix and Seraseq™ Myeloid Fusion RNA Mix. These well-characterized positive reference materials contain the most important genetic alterations across many different types of myeloid cancers, and can ultimately help increase scientists’ and clinicians’ confidence in assay results and patient reporting.
If you're looking for the best QC solution for your NGS myeloid cancer test, and would like to learn more click here. You can also contact us via the comments below.
References
- Barbui T et al. “Rationale for revision and proposed changes of the WHO diagnostic criteria for polycythemia vera, essential thrombocythemia and primary myelofibrosis.” Blood Cancer J. 2015 Aug 5(8): e337, doi: 10.1038/bcj.2015.64. Accessed 3 Oct 2017.
- Gelsi-Boyer V et al. “Mutations in ASXL1 are associated with poor prognosis across the spectrum of malignant myeloid diseases.” Journal of Hematology & Oncology 2012 5:12, https://doi.org/10.1186/1756-8722-5-12. Accessed 3 Oct 2017.
- Larrosa-Garcia M et al. “FLT3 Inhibitors in Acute Myeloid Leukemia: Current Status and Future Directions.” Molecular Cancer Therapeutics 2017 16(6);991-1001, doi: 10.1158/1535-7163.MCT-16-0876. Accessed 3 Oct 2017.
- FDA drug label, RYDAPT®: https://www.accessdata.fda.gov/drugsatfda_docs/label/2017/207997s000lbl.pdf
- Mondesir J et al. “IDH1 and IDH2 mutations as novel therapeutic targets: current perspectives.” J Blood Med 2016 7:171-180, doi: 10.2147/JBM.S70716. Accessed 3 Oct 2017.
- FDA drug label, IDHIFA®: https://www.accessdata.fda.gov/drugsatfda_docs/label/2017/209606s000lbl.pdf
- Falini B et al. “BRAF-V600E mutation in hairy cell leukemia: from bench to bedside.” Blood 2016 prepublished online Aug 23, 2016, doi: 10.1182/blood-2016-07-418434. Accessed 3 Oct 2017.





