Choose your Article Focus | NGS | Molecular & Serology
Development of a Novel Reference Material for MRD Monitoring Assays
Category: MRD, ccfDNA, cfDNA, Minimal Residual Disease, NGS, ctDNA, reference materials
Posted by
Yves Konigshofer, PhD on Aug 17, 2021 12:00:00 AM
Authors: Yves Konigshofer, PhD; Andrew Anfora, PhD; Omo Clement, PhD; and Krystyna Nahlik, PhD. LGC Clinical Diagnostics. Introduction Liquid biopsy methods developed for circulating tumor DNA (ctDNA) analysis in solid tumors are transforming clinical practice, allowing for non-invasive detection and assessment of earlier stages of disease, monitoring for resistance to therapy, and post-treatment monitoring for minimal residual disease (MRD) and recurrence of cancer. The presence of minimal residual disease may be prognostic in that is has been found to correlate with worse patient outcomes, so early and accurate measurement is crucial. ctDNA-based assays allow for the detection of MRD at earlier time points than standard clinical and imaging surveillance, and could allow for treatment modification based on real-time assessment of the tumor genomic landscape.
0 Comments Click here to read/write comments
How to resolve the challenges of MRD?
Category: MRD, cfDNA, NGS, ctDNA, reference materials
Posted by
Yves Konigshofer, PhD on Oct 7, 2020 12:00:00 AM
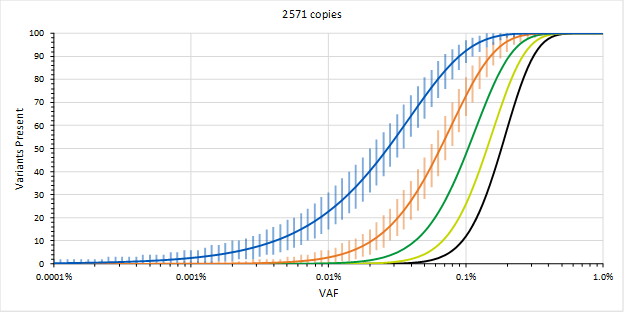
This is part 2 of 2 of the MRD blog post. (Click here for part 1). In this section, we will discuss how to overcome some of the most common challenges of MRD testing. Overcoming the Challenges In order to mitigate sequencing errors, methods using Molecular Barcodes (MBCs), Unique Molecular Identifiers (UMIs), etc. (which are essentially all the same) may be used, where each starting molecule is sequenced many times. The MBCs are then used to generate consensus sequences from sequences that were likely obtained from the same starting molecule. The assumption is that errors appear due to somewhat stochastic processes and that the consensus sequences will likely be correct. This requires many observations of the same starting molecule, so it will be recommended to generate 10-fold more sequences than there are molecules. Therefore, with 8,000 genomic equivalents, we might want to target a sequencing depth of 80,000. This is a reason why using 10-fold more input ccfDNA may not necessarily be a good thing (in addition to having to obtain a 10-fold larger liquid biopsy) since we may have to increase sequencing depth accordingly to 800,000, which could increase the cost of sequencing 10-fold, which could reduce the likelihood for payment and running the assay profitably.
0 Comments Click here to read/write comments
So, you want to monitor Measurable Residual Disease? What are the challenges?
Category: MRD, cfDNA, Minimal Residual Disease, NGS, ctDNA, reference materials
Posted by
Yves Konigshofer, PhD on Oct 1, 2020 12:00:00 AM
Part 1 of 2 Background Measurable Residual Disease (MRD) monitoring – for purposes of this blog – will be the act of looking for somatic variants in a liquid biopsy sample by analyzing circulating cell-free DNA (ccfDNA). This is done to monitor the disappearance of a metastatic solid tumor during treatment and to follow any future reemergence of that cancer. Analyzing ccfDNA assumes that circulating tumor DNA (ctDNA) will be present, and the median ctDNA frequency in patient samples across cancers seems to be around 0.5 to 1 %. Thus, the median variant allele frequencies (VAFs) of the somatic variants will start around this range, and the goal of MRD monitoring is to be able to detect them at much lower VAFs. This can be challenging and if we are going to design an assay for MRD monitoring, then we need to be aware of them and overcome them.
0 Comments Click here to read/write comments