During my time spent in academia, I studied in a laboratory focused on molecular switches. Although cellular signaling cascades appear as nothing more than incomprehensible spiderwebs when viewed graphically, the concept is very straightforward: certain molecules can act to regulate cellular processes by functioning like electrical switches. Just as light switches in your home complete or break a circuit to control the flow of electrical current, molecular switches such as Ras GTPases (PDF) control whether certain signals can elicit a cellular response.

Pathways that regulate cell growth, differentiation, and survival are of particular interest for clinical oncology. Unsurprisingly, just as a broken light switch can make it difficult to turn off a light, mutations that disrupt signaling such as the epidermal growth factor receptor (EGFR) pathway can lead to tumor formation. To learn more about the larger picture of oncogenesis, a seminal publication by Douglas Hanahan and Robert Weinberg reviews characteristics necessary for the formation and progression of tumors which the authors refer to as the “Hallmarks of Cancer” – Part I was published in Cell in 2000 (PDF), and an update followed in 2011 (PDF). I also had the privilege of hearing Dr. Hanahan discuss the hallmarks of cancer in the context of advances in precision medicine at the American Association of Cancer Research 2016 meeting. Of particular interest from this talk was the observation that therapies targeted to single hallmarks alone have proven largely ineffective, and that combination therapies which target multiple hallmarks simultaneously may be a necessity.
Understanding these signaling pathways in great depth has tremendous clinical value for identification of promising therapeutic targets. To continue with the EGFR pathway as an example, inhibition of EGFR with different classes of molecules has become central to cancer chemotherapy. Two main classes of drugs are currently used to down-regulate EGFR signaling:
- Monoclonal antibody inhibitors function as receptor antagonists by preventing the epidermal growth factor ligand from binding the receptor, thus reducing cellular response to EGFR signaling.
- Small-molecule kinase inhibitors act on the domain of the EGFR molecule that is located on the interior of the cell membrane to prevent signaling to downstream effector proteins.
Ideally, down-regulation of a pathway that promotes cell growth and survival should halt or slow tumor progression. However, resistance to anti-EGFR therapies inevitably occurs over the course of treatment, and is a textbook example of the ongoing arms race between cancer therapeutics and complex, highly heterogeneous malignancies.
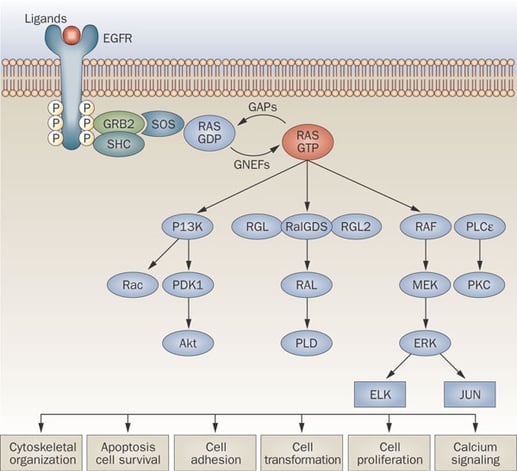
Image courtesy of Normanno et al. 2009. Implications for KRAS status and EGFR-targeted therapies in metastatic CRC. Nat Rev Clin Onc. 6:519-527.
It is logical to suspect that one cause of insensitivity to EGFR inhibition may be disruption of a downstream signaling molecule such that the pathway remains hyper-activated irrespective of the receptor itself. Indeed, it has been discovered that mutations in KRAS, a Ras GTPase which acts as a key molecular switch in the EGFR signaling pathway, can confer resistance to anti-EGFR therapies. Therefore, for patients affected by tumors carrying pathogenic KRAS mutations, therapeutic approaches outside of EGFR inhibitors must be taken to avoid wasted time and increased cost that ineffective therapies would incur.
Since the discovery of this chemotherapeutic resistance mechanism, the KRAS oncogene has become an essential diagnostic target for cancer patients potentially eligible to receive anti-EGFR treatment. Various KRAS molecular assays have been approved by the FDA as companion diagnostic (CDx) tests for EGFR inhibitors, among them the cobas® KRAS Mutation Test from Roche Molecular Systems, Inc. (PDF) and the therascreen® KRAS RGQ PCR Kit from QIAGEN Manchester, Ltd (PDF). These and other CDx tests are regarded by the Agency as Class III medical devices, which is the highest risk category which requires very rigorous validation in order to receive approval. For clinical laboratories running KRAS assays, it is vital to ensure that these tests are performing well, especially when pushed to the limits of their sensitivity.
In order to assist clinical laboratories as well as diagnostic manufacturers who are running or developing highly-targeted KRAS molecular assays for solid tumor specimens, SeraCare has developed the Seraseq™ FFPE Tumor KRAS Reference Material Kit. This new reference material contains 7 pathogenic KRAS mutations that account for approximately 97% of characterized mutation subtypes. Cells containing KRAS mutations are blended with a well-characterized wild-type background to achieve Variant Allele Frequencies around the Limit of Detection for industry-leading KRAS diagnostic assays, and are orthogonally tested using qualified digital PCR assays. For each of the 7 mutations, one 5-micron formalin-fixed paraffin-embedded section is produced for consistency with the cobas KRAS Mutation Test and therascreen KRAS RGQ PCR Kit instructions for use, which require 5-µm FFPE sections. The 7 FFPE sections are packed into separate vials, and the set of 7 vials is provided in a convenient kit format.
To learn more about this product, as well as other oncology reference materials from SeraCare, please visit us online or contact us here or via the comments below.




