Overview
Tumor mutational burden (TMB) is a measurement of the number of mutations carried by a tumor cell genome. By comparing DNA sequences from a patient’s healthy tissue and tumor cells, and using a number of complex algorithms, scientists can determine the number of acquired somatic mutations present in tumors but not in normal tissues.1 NGS is the primary method employed to measure TMB, either through targeted panels or whole exome sequencing (WES);2 the latter is considered the gold-standard measurement today.
Clinical application of TMB for I-O treatment decision however faces current challenges, such as (a) lack of standardization in TMB measurements across different NGS panels, and (b) lack of qualified reference standards to provide ground-truth data in harmonizing these measurements. To address these challenges, advocacy organizations in the US and EU are coordinating with industry stake holders to align TMB scores observed across the different TMB targeted NGS panels. These initiatives are critical to ensuring a wider access to analytically validated NGS-based assays as clinical support tools to cancer patients, cancer treatment hospitals, as well as drug manufacturers.
LGC SeraCare is a member of the TMB Harmonization Project led by Friends of Cancer Research (FoCR)3 consortium working to harmonize the variability in TMB measurements across different targeted NGS panels from those derived using the gold standard whole exome sequencing (WES). In this role, LGC SeraCare was the sole provider of TMB reference standards to participating members,4 becoming industry’s first supplier of TMB assay reference materials.5 Additionally, LGC SeraCare partnered with top industry experts in NGS assay development and bioinformatics6 to build the most comprehensively validated TMB reference standards.
Development and Characterization of TMB Reference Standards
Genomic DNA and FFPE formats of SeraCare’s TMB reference materials were formulated as described in Figure 1, from human diseased cell lines and their matched normal cell lines, in either a 100% tumor-normal set (gDNA TMB controls) or 30% tumor blend (FFPE TMB controls).7 In Figure 2 is the outline of the bioinformatics analysis pipeline for TMB score determination, and in Figure 3 are results of a multi laboratory evaluation of these materials using targeted NGS panels from laboratories of members in the LGC SeraCare TMB Working Group.6
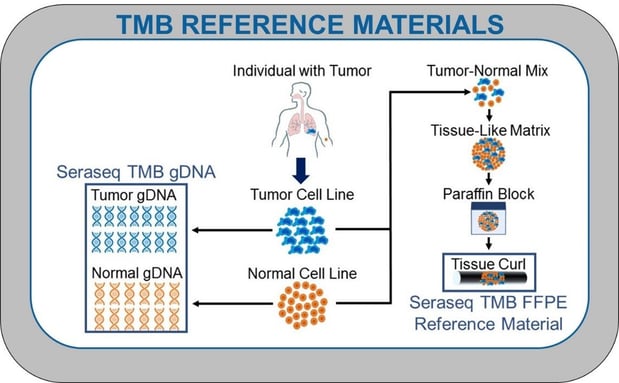
Figure-1: Simplified view of the TMB reference material (gDNA and FFPE) development process.7
TMB Measurements by Whole Exome Sequencing (WES)
Whole exome sequencing analysis of extracted gDNA from the tumor and normal cell lines were performed using the Agilent SureSelect Exome enrichment kit sequenced on an ILMN Novaseq 6000 NGS system. Typical sequencing metrics for the tumor and normal samples are as shown in Table 1 below. Variant analysis (Figure 2) and TMB scoring were deduced as described by FOCR’s Phase I project recommendations.9
Table 1: Representative whole exome sequencing (WES) metrics for tumor and normal TMB reference materials.
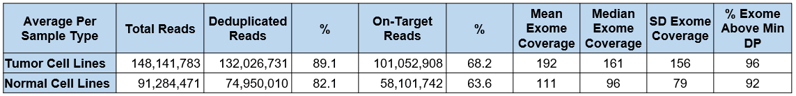
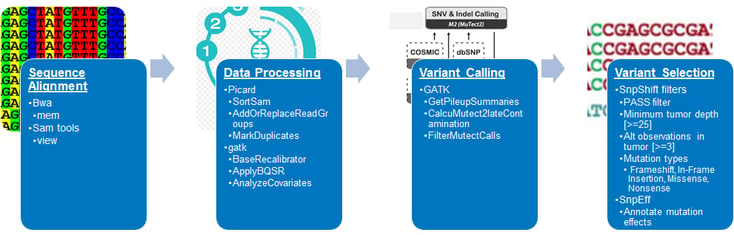
Figure 2: SeraCare TMB analysis pipeline7 based on FoCR TMB harmonization project recommendation.9
TMB Scores by WES and Annotated Scores derived using Targeted NGS Panels
Table-2: WES-determined TMB scores (non-synonymous mutations) for Seraseq TMB reference materials (gDNA & FFPE)
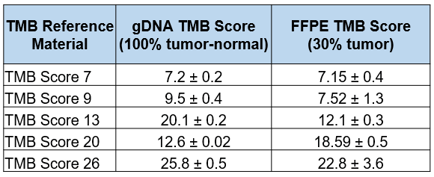
Table-3: Targeted NGS panel-generated TMB scores (non-synonymous mutations) for Seraseq TMB reference materials.6
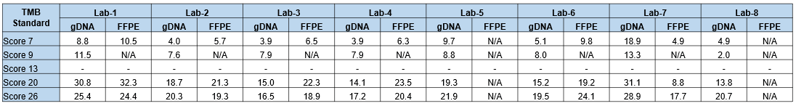
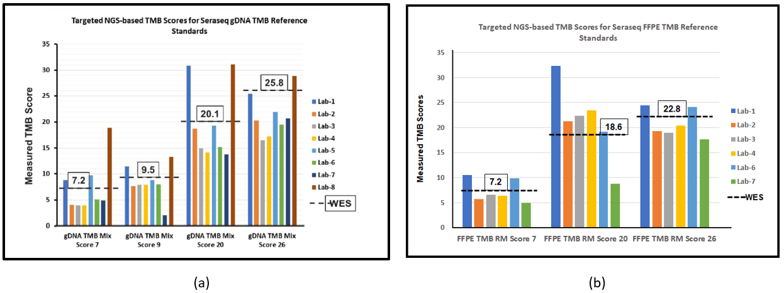
Figure 3: Multi-laboratory targeted NGS panel evaluation6 and measurements of TMB scores for the (a) Seraseq gDNA and (b) FFPE TMB reference materials. Boxed TMB scores represent the corresponding whole exome sequencing (WES) scores.7
Seraseq® TMB Products
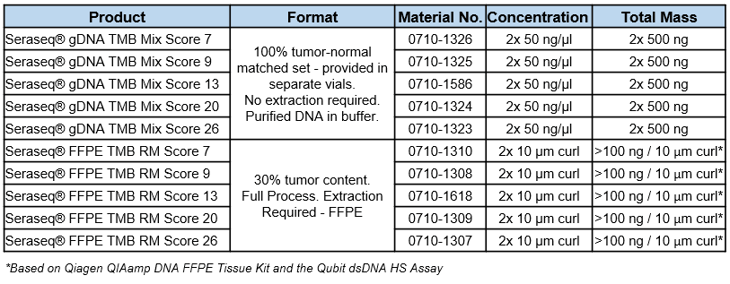
References
- https://www.scientificamerican.com/custom-media/tumor-mutation-burden/
- https://www.decibio.com/218/03/13/tumor-mutation-burden-tmb-immuno-oncology/
- https://www.focr.org/
- https://www.focr.org/news/friends-cancer-research-announces-launch-phase-ii-tmb-harmonization-project
- TMB Reference Standard Blog post
- SeraCare TMB Working Group consists of participating members from the following organizations: Illumina (ILMN), Astra Zeneca (AZ), National Institute of Standards and Technology (NIST), Asuragen Inc., Moffit Cancer Center (Moffit), The Broad Institute, Neo New Oncology GmbH, InterMountain Precision Cancer Care (Navican), Sema4 Genomics (Sema4), Roche Sequencing (Roche), Medical College of Augusta University (MCG), and Memorial Sloan Kettering Cancer Center (MSKCC).
- (a) Butler, Matt, et al, Characterization of tumor-normal cell line pairs for TMB Standardization. 2019 AMP Meeting, Poster #ST084; (b) Butler, M.B., et. al., Development of Reference Materials for NGS-based Tumor Mutational Burden (TMB) Measurement and Standardization, Cancer Genomics 2019 EACR Conference, Cambridge, UK; (c) Butler, M.B., et. al., Improving and Standardizing TMB Assay Performance, 2019 AACR Meeting, Atlanta, GA, USA.
- MoCha = Molecular Characterization Laboratory, Frederick National Lab for Cancer Research (NCI)
- Merino, D., et. al., 2019 ASCO Meeting, Chicago, USA, Poster #268; https://meetinglibrary.asco.org/record/172797/abstract
- Kohle, R., et. al., Guideline adherent clinical validation of a comprehensive DNA/RNA panel (523-genes TruSight Oncology 500) for determination of single nucleotide variants (SNVs) small insertions or deletions (Indels), copy number variations (CNVs), splice variations (SVs), gene fusions (GFs), tumor mutation burden (TMB) and microsatellite instability (MSI) on a next generation sequencing platform in a CLIA setting, 2019 AACR Meeting, Atlanta, GA, USA.




