Of the many fantastic posters presented at AMP’s Annual Meeting in San Antonio, two concerning NGS-based liquid biopsy assays stood out. Both presenters described how their organizations are working to reliably detect pathogenic variants at extremely low allele frequencies – efforts critical to the clinical adoption of NGS-based liquid biopsy assays.
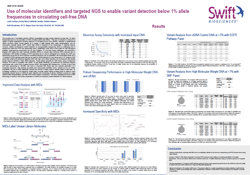
Swift Biosciences presented some very interesting data on molecular identifiers (MIDs) that enable accurate variant detection below 1% AF in ctDNA. They used the Seraseq ctDNA Reference Material v2 to test their assay’s limit of detection. Click here to download the poster and watch Swift’s Francesco Criscuolo explain how Swift’s MID technology can improve NGS data analysis with his poster Use of Molecular Identifiers and Targeted NGS to Enable Variant Detection Below 1% Allele Frequencies in Circulating Cell-Free DNA.

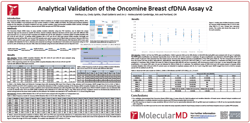
Another compelling poster was presented by MolecularMD’s Dr. WeiHua Liu. Her team took a methodical approach to validating the Oncomine Breast cfDNA Assay v2 and used both the Seraseq ctDNA Complete Mutation Mix and ctDNA Mutation Mix v2 reference materials to determine the amount of DNA input needed to reproducibly detect SNVs at 0.1% AF. Check out her poster presentation video, titled Analytical Validation of Oncomine Breast cfDNA Assay v2, to learn more about their methodology.
Want to learn more about our QC and validation solutions for NGS-based liquid biopsy assays? Click here.
To watch the videos and download these free posters, click each title below: